File Formats
The fasta format was invented in 1988 and designed to represent nucleotide or peptide sequences. It originates from the FASTA software package, but is now a standard in the world of bioinformatics.
The first line in a FASTA file starts with a ">" (greater-than) symbol followed by the description or identifier of the sequence. Following the initial line (used for a unique description of the sequence) is the actual sequence itself in standard one-letter code.
A few sample sequences:
>KX580312.1 Homo sapiens truncated breast cancer 1 (BRCA1) gene, exon 15 and partial cds
GTCATCCCCTTCTAAATGCCCATCATTAGATGATAGGTGGTACATGCACAGTTGCTCTGGGAGTCTTCAG
AATAGAAACTACCCATCTCAAGAGGAGCTCATTAAGGTTGTTGATGTGGAGGAGTAACAGCTGGAAGAGT
CTGGGCCACACGATTTGACGGAAACATCTTACTTGCCAAGGCAAGATCTAG
>KRN06561.1 heat shock [Lactobacillus sucicola DSM 21376 = JCM 15457]
MSLVMANELTNRFNNWMKQDDFFGNLGRSFFDLDNSVNRALKTDVKETDKAYEVRIDVPGIDKKDITVDY
HDGVLSVNAKRDSFNDESDSEGNVIASERSYGRFARQYSLPNVDESGIKAKCEDGVLKLTLPKLAEEKIN
GNHIEIE
A fasta file can contain multiple sequence. Each sequence will be separated by their "header" line, starting by ">".
Example:
>KRN06561.1 heat shock [Lactobacillus sucicola DSM 21376 = JCM 15457]
MSLVMANELTNRFNNWMKQDDFFGNLGRSFFDLDNSVNRALKTDVKETDKAYEVRIDVPGIDKKDITVDY
HDGVLSVNAKRDSFNDESDSEGNVIASERSYGRFARQYSLPNVDESGIKAKCEDGVLKLTLPKLAEEKIN
GNHIEIE
>3HHU_A Chain A, Human Heat-Shock Protein 90 (Hsp90)
MPEETQTQDQPMEEEEVETFAFQAEIAQLMSLIINTFYSNKEIFLRELISNSSDALDKIRYESLTDPSKL
DSGKELHINLIPNKQDRTLTIVDTGIGMTKADLINNLGTIAKSGTKAFMEALQAGADISMIGQFGVGFYS
AYLVAEKVTVITKHNDDEQYAWESSAGGSFTVRTDTGEPMGRGTKVILHLKEDQTEYLEERRIKEIVKKH
SQFIGYPITLFVEK
The fastq format is also a text based format to represent nucleotide sequences, but also contains the corresponding quality of each nucleotide. It is the standard for storing the output of high-throughput sequencing instruments such as the Illumina machines.
A fastq file uses four lines per sequence:
· Line 1 begins with a '@' character and is followed by a sequence identifier and an optional description (like a FASTA title line).
· Line 2 is the raw sequence letters.
· Line 3 begins with a '+' character and is optionally followed by the same sequence identifier (and any description) again.
· Line 4 encodes the quality values for the sequence in Line 2, and must contain the same number of symbols as letters in the sequence.
An example sequence in fastq format:
@SEQ_ID
GATTTGGGGTTCAAAGCAGTATCGATCAAATAGTAAATCCATTTGTTCAACTCACAGTTT
+
!''*((((***+))%%%++)(%%%%).1***-+*''))**55CCF>>>>>>CCCCCCC65
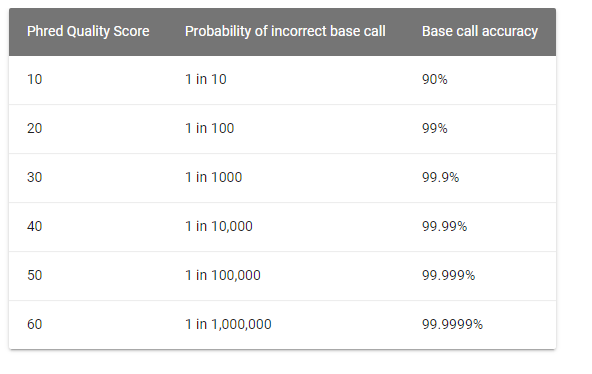
The quality, also called phred score, is the probability that the corresponding basecall is incorrect.
Phred scores use a logarithmic scale, and are represented by ASCII characters, mapping to a quality usually going from 0 to 40.

From Wikipedia:
SAM (file format) is a text-based format for storing biological sequences aligned to a reference sequence developed by Heng Li. The acronym SAM stands for Sequence Alignment/Map. It is widely used for storing data, such as nucleotide sequences, generated by Next generation sequencing technologies and usually mapped to a reference.
The SAM format consists of a header and an alignment section. The binary representation of a SAM file is a BAM file, which is a compressed SAM file.[1] SAM files can be analysed and edited with the software SAMtools.
The SAM format has a really extensive and complex specification that you can find here.
In brief it consists of a header section and reads (with other information) in tab delimited format.
@HD VN:1.0 SO:unsorted
@SQ SN:O_volvulusOVOC_OM1a LN:2816604
@SQ SN:O_volvulusOVOC_OM1b LN:28345163
@SQ SN:O_volvulusOVOC_OM2 LN:25485961
M01137:130:00-A:17009:1352/14 * 0 0 * * 0 0 AGCAAAATACAACGATCTGGATGGTAGCATTAGCGATGCGACACTGCTTGAACCGTCAAAG FGGFGCFGFFGC8,,@D?E6EFCF,=AEFFGGDGGGADFGG@>FFEGGG:+<7D>AFCFGG YT:Z:UU
The vcf format is also a text-based file format. VCF stands for Variant Call Format and is used to store gene sequence variations (SNVs, indels). The format has been developped for genotyping projects, and is the standard to represent variations in the genome of a species.
A vcf is a tab-delimited file, described here.
##fileformat=VCFv4.0
##fileDate=20110705
##reference=1000GenomesPilot-NCBI37
##phasing=partial
##INFO=<ID=NS,Number=1,Type=Integer,Description="Number of Samples With Data">
##INFO=<ID=DP,Number=1,Type=Integer,Description="Total Depth">
##INFO=<ID=AF,Number=.,Type=Float,Description="Allele Frequency">
##INFO=<ID=AA,Number=1,Type=String,Description="Ancestral Allele">
##INFO=<ID=DB,Number=0,Type=Flag,Description="dbSNP membership, build 129">
##INFO=<ID=H2,Number=0,Type=Flag,Description="HapMap2 membership">
##FILTER=<ID=q10,Description="Quality below 10">
##FILTER=<ID=s50,Description="Less than 50% of samples have data">
##FORMAT=<ID=GQ,Number=1,Type=Integer,Description="Genotype Quality">
##FORMAT=<ID=GT,Number=1,Type=String,Description="Genotype">
##FORMAT=<ID=DP,Number=1,Type=Integer,Description="Read Depth">
##FORMAT=<ID=HQ,Number=2,Type=Integer,Description="Haplotype Quality">
#CHROM POS ID REF ALT QUAL FILTER INFO FORMAT Sample1 Sample2 Sample3
2 4370 rs6057 G A 29 . NS=2;DP=13;AF=0.5;DB;H2 GT:GQ:DP:HQ 0|0:48:1:52,51 1|0:48:8:51,51 1/1:43:5:.,.
2 7330 . T A 3 q10 NS=5;DP=12;AF=0.017 GT:GQ:DP:HQ 0|0:46:3:58,50 0|1:3:5:65,3 0/0:41:3
2 110696 rs6055 A G,T 67 PASS NS=2;DP=10;AF=0.333,0.667;AA=T;DB GT:GQ:DP:HQ 1|2:21:6:23,27 2|1:2:0:18,2 2/2:35:4
2 130237 . T . 47 . NS=2;DP=16;AA=T GT:GQ:DP:HQ 0|0:54:7:56,60 0|0:48:4:56,51 0/0:61:2
2 134567 microsat1 GTCT G,GTACT 50 PASS NS=2;DP=9;AA=G GT:GQ:DP 0/1:35:4 0/2:17:2 1/1:40:3
chr1 45796269 . G C
chr1 45797505 . C G
chr1 45798555 . T C
chr1 45798901 . C T
chr1 45805566 . G C
chr2 47703379 . C T
chr2 48010488 . G A
chr2 48030838 . A T
chr2 48032875 . CTAT -
chr2 48032937 . T C
chr2 48033273 . TTTTTGTTTTAATTCCT -
chr2 48033551 . C G
chr2 48033910 . A T
chr2 215632048 . G T
chr2 215632125 . TT -
chr2 215632155 . T C
chr2 215632192 . G A
chr2 215632255 . CA TG
chr2 215634055 . C T
The general feature format (gff) is another text file format, used for describing genes and other features of DNA, RNA and protein sequences. It is the standard for annotation of genomes.
A gff file should contain 9 columns, described here
##description: evidence-based annotation of the human genome (GRCh38), version 25 (Ensembl 85)
##provider: GENCODE
##contact: gencode-help@sanger.ac.uk
##format: gtf
##date: 2016-07-15
chr1 HAVANA gene 11869 14409 . + . gene_id "ENSG00000223972.5"; gene_type "transcribed_unprocessed_pseudogene"; gene_status "KNOWN"; gene_name "DDX11L1"; level 2; havana_gene "OTTHUMG00000000961.2";
chr1 HAVANA transcript 11869 14409 . + . gene_id "ENSG00000223972.5"; transcript_id "ENST00000456328.2"; gene_type "transcribed_unprocessed_pseudogene"; gene_status "KNOWN"; gene_name "DDX11L1"; transcript_type "processed_transcript"; transcript_status "KNOWN"; transcript_name "DDX11L1-002"; level 2; transcript_support_level "1"; tag "basic"; havana_gene "OTTHUMG00000000961.2"; havana_transcript "OTTHUMT00000362751.1";
chr1 HAVANA exon 11869 12227 . + . gene_id "ENSG00000223972.5"; transcript_id "ENST00000456328.2"; gene_type "transcribed_unprocessed_pseudogene"; gene_status "KNOWN"; gene_name "DDX11L1"; transcript_type "processed_transcript"; transcript_status "KNOWN"; transcript_name "DDX11L1-002"; exon_number 1; exon_id "ENSE00002234944.1"; level 2; transcript_support_level "1"; tag "basic"; havana_gene "OTTHUMG00000000961.2"; havana_transcript "OTTHUMT00000362751.1";
chr1 HAVANA exon 12613 12721 . + . gene_id "ENSG00000223972.5"; transcript_id "ENST00000456328.2"; gene_type "transcribed_unprocessed_pseudogene"; gene_status "KNOWN"; gene_name "DDX11L1"; transcript_type "processed_transcript"; transcript_status "KNOWN"; transcript_name "DDX11L1-002"; exon_number 2; exon_id "ENSE00003582793.1"; level 2; transcript_support_level "1"; tag "basic"; havana_gene "OTTHUMG00000000961.2"; havana_transcript "OTTHUMT00000362751.1";
chr1 HAVANA exon 13221 14409 . + . gene_id "ENSG00000223972.5"; transcript_id "ENST00000456328.2"; gene_type "transcribed_unprocessed_pseudogene"; gene_status "KNOWN"; gene_name "DDX11L1"; transcript_type "processed_transcript"; transcript_status "KNOWN"; transcript_name "DDX11L1-002"; exon_number 3; exon_id "ENSE00002312635.1"; level 2; transcript_support_level "1"; tag "basic"; havana_gene "OTTHUMG00000000961.2"; havana_transcript "OTTHUMT00000362751.1";